The hisat program can automatically download SRA data as needed. In some cases, users may want to download SRA data and retain a copy. To download using NCBI's 'prefetch' tool, you would need to set up your own configuration file for the NCBI SRA toolkit. Use the command vdb-config to set up a directory for downloading. SRA toolkit contains important tools to manipulate SRA (Short Read Archive) file. The objective of this article is to show you, how to install SRA toolkit on Ubuntu/Linux system. Download the last version for your computer operating system from here Use the following command on Linux to download the file sratoolkit.2.4.1.
Using SRAtoolkit
SRA toolkit has been configured to connect to NCBI SRA and download via FTP. The simple command to fetch a SRA file you can use this command:
This will download the SRA file (in
sra format) and then convert them to fastq file for you.If your SRA file is paired, you will still end up with a single fastq file, since, fastq-dump, by default writes them as interleaved file. To change this, you can provide --split-files argument.The downloaded fastq files will have
sra number suffixed on all header lines of fastq fileAlthough, this normally does not affect any programs, some programs might throw an error saying that it can’t process these
fastq files. To avoid this, you an request the file to be in the orignal format (--origfmt). Also, note that if you’re downloading files in bulk, you can save a lot of space by compressing them in gzip format (--gzip).The
fastq-dump is also capable of doing:- Additional filtering or clipping of the downloaded reads: to remove reads with poor quality or to trim adapters. Although, this will work for the single end reads, for paired-end reads it may cause differential treatment for each pairs and might not be usable for mapping programs that needs strict pairs.
- Compressed format: either as gzipped or bzipped files using
--gzipor--bzip2options. - fasta format: by using the
--fastaoption
Using Linux commands:
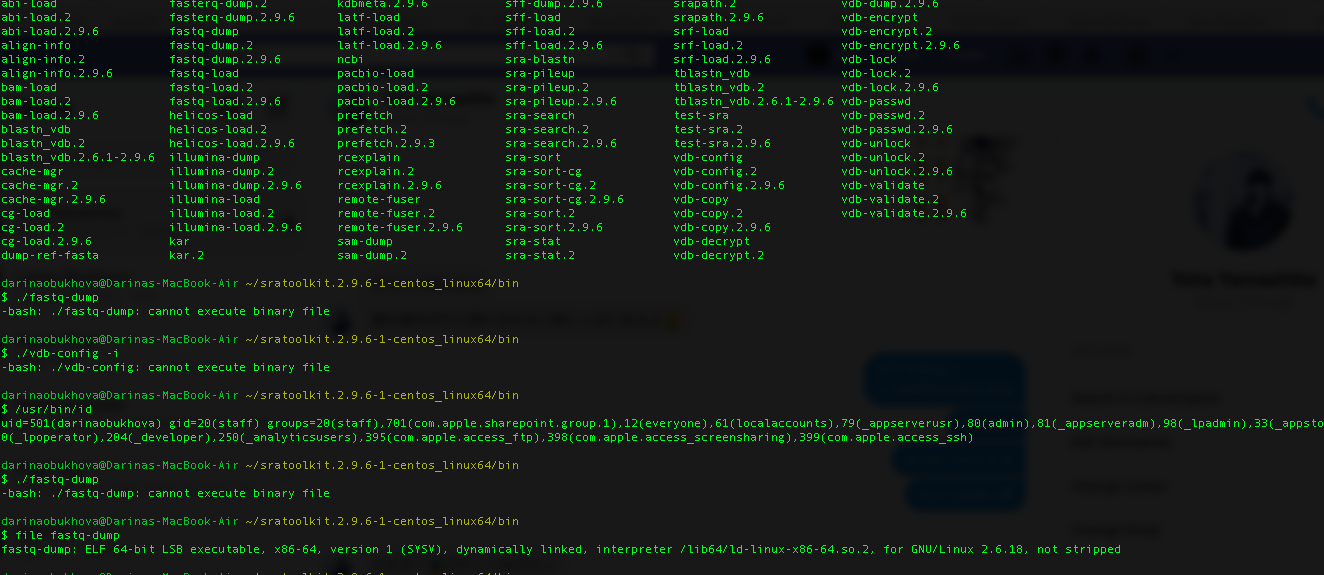
In cases were you cannot run the SRA toolkit or any other programs to download the file, you can still use the inbuilt commands of Linux such as
wget and curl. The standard web link for downloading the SRA files is:You need to replace the
SRRNNNNNN with the actual SRR number for it to work.You can either use
wgetor
curlIf you have a large list of ids, you can simply loop it over using a
while loopThe datasets can also be downloaded from DDBJ or EMBL using the FTP links, but the transfer speeds might be affected if you’re not near their servers.
Download Sra Toolkit Mac Terminal Linux
Using Aspera Connect (ascp)

Sra Toolkit Manual
Aspera uses high-speed file transfer to rapidly transfer large files and data sets over an existing WAN infrastructure.
To get the
sra files:This usually prefetches the SRA file to your home directory in folder named ncbi. If your home directory does not contain enough space to store all data, you may want to create another directory and softlink to the home. To do this:
when you run this, you will have a directory named
ncbi in your home, but the data is actually stored in /project/storage/your_dir/ncbiThen you can convert the SRA files back to fastq format using
fastq-dump command.Downloading all SRA files related to a BioProject/study
NCBI Sequence Read Archive (SRA) stores sequence and quality data (fastq files) in aligned or unaligned formats from NextGen sequencing platforms. A BioProject is a collection of biological data related to a single initiative, originating from a single organization or from a consortium. A BioProject record provides users a single place to find links to the diverse data types generated for that project. Often times, once single BioProject will hold a considerable number of experiments and it gets tedious to download them all individually. Here is the guide to show how to do this in a effecient way:
First load the modules that are needed:
To get the SRR numbers associated with the project:
To download them all in parallel (limit the number to 3 concurrent downloads)
How To Use Sra Toolkit
Make sure you do this on Condoddtn node or as a PBS job
